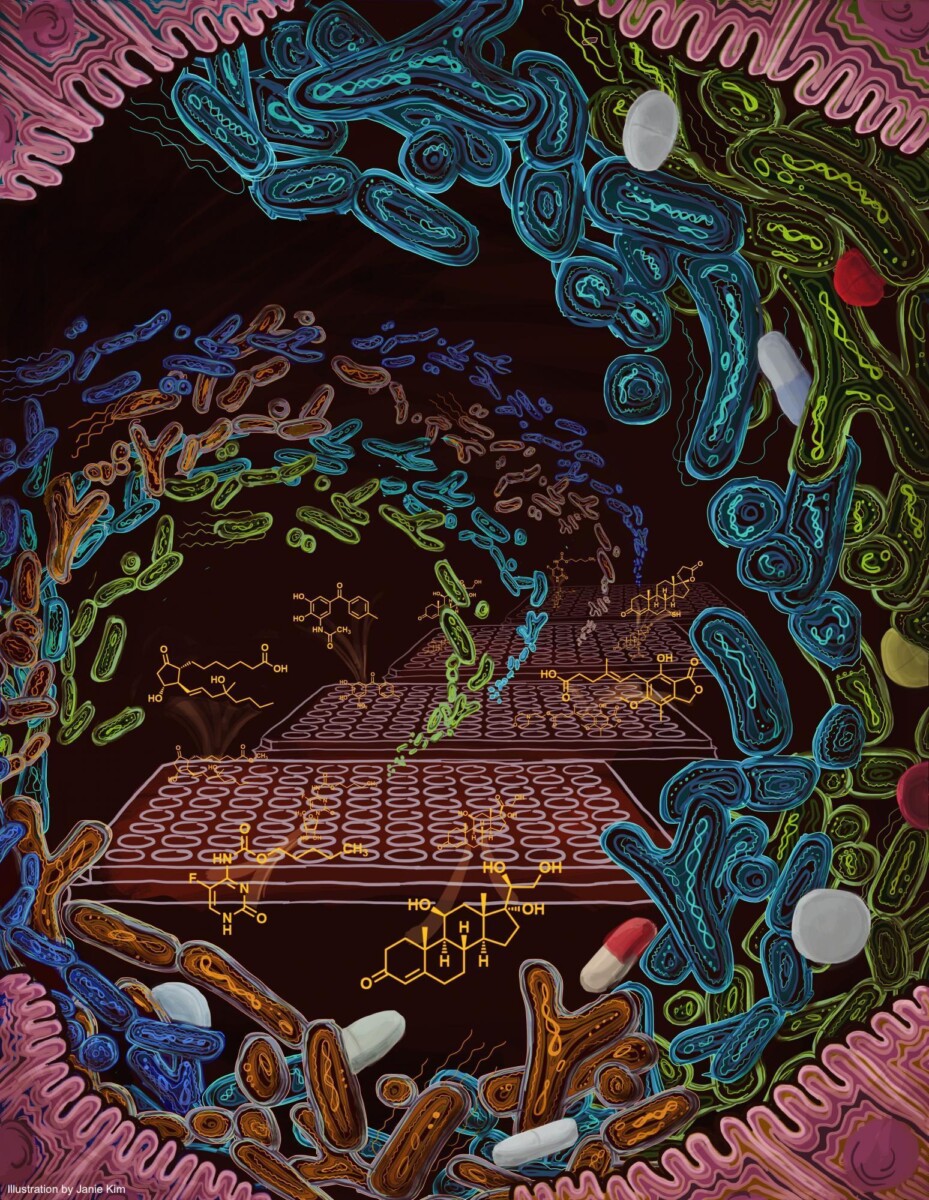

In addition to the de novo production of small molecules, the human gut microbiome encodes a large repertoire of biochemical enzymes that can metabolize exogenous small molecules – whether they are derived from the human host, or from dietary or therapeutic sources. This concept has been explored for decades by using single isolates of the human microbiome against a common set of human-, drug-, or diet-derived chemicals, but has not been explored systematically. We aim to expand this area of research by developing high-throughout, unbiased approaches for defining the space of biochemical transformations that can be exerted by the collective human gut microbiome against a wide variety of chemicals, and the pharmacological consequences of these transformations.
In the case of marine organisms (e.g., sponges, tunicates, mollusks, algae) and their symbionts, small-molecule-mediated interactions can provide the host with indispensable means of chemical defense, allowing it to survive in a predator-rich environment. We characterize these interactions by first determining the true microbial producers of the defensive molecules and the molecular mechanisms of their production, then we study the evolution of the symbiosis in light of the production of these molecules, and finally, we study the ecological consequences of their production for all interacting partners.
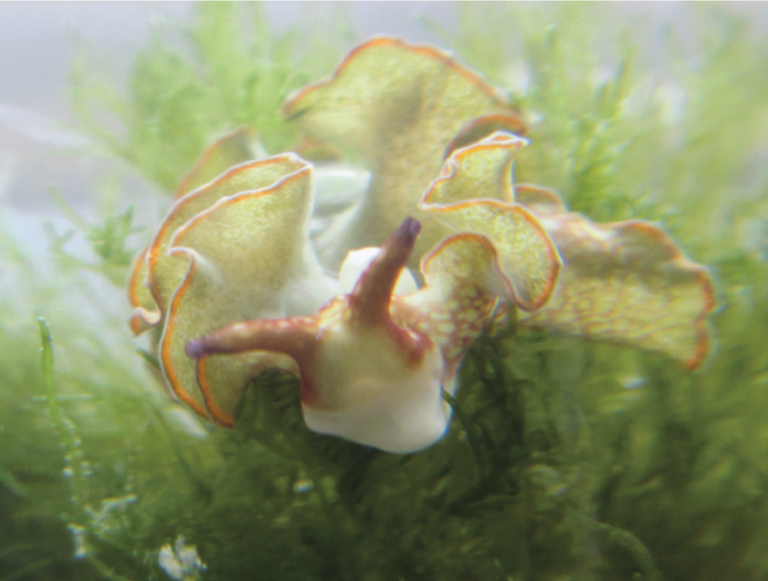
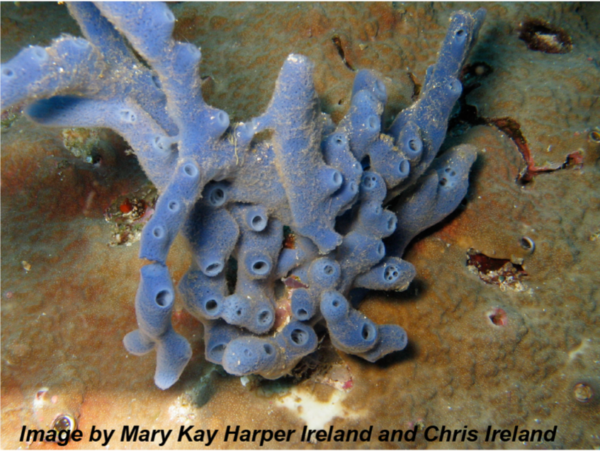


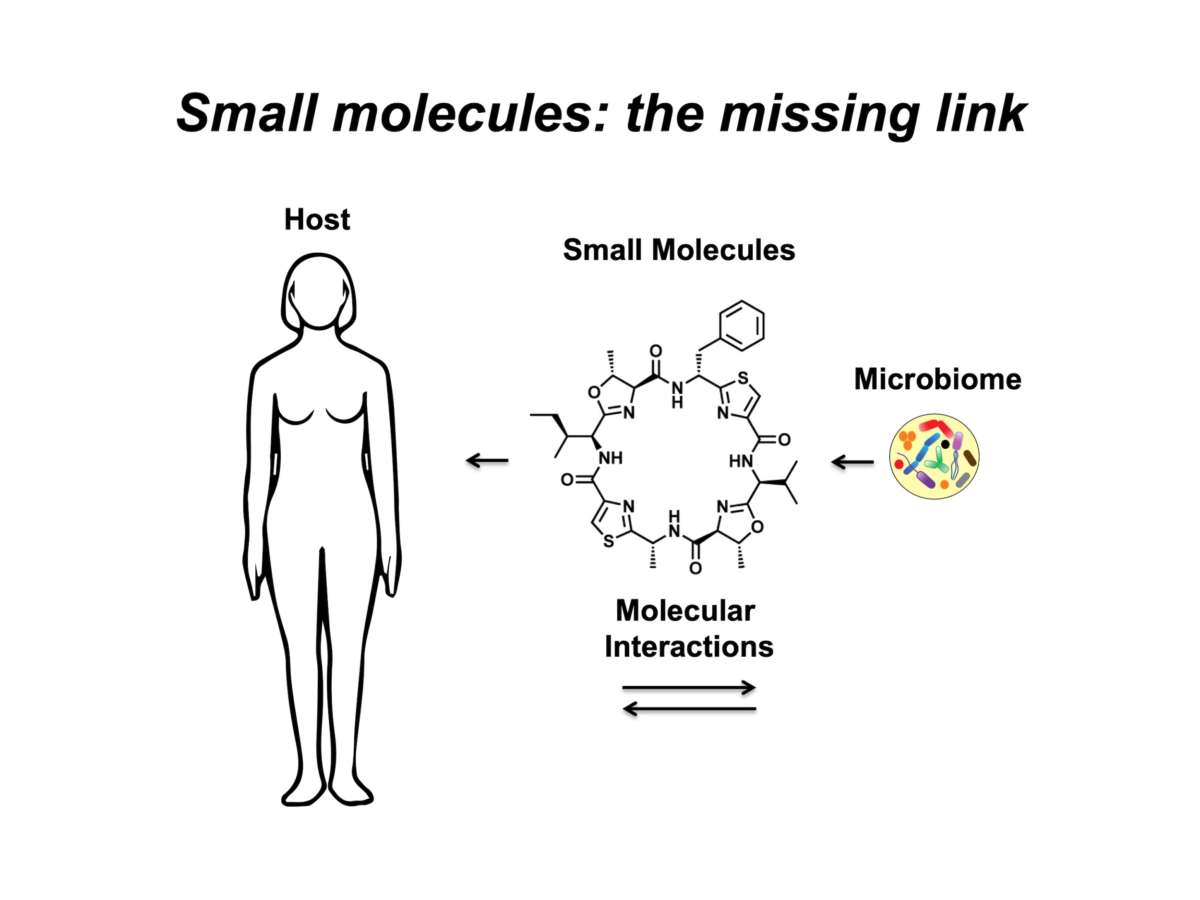
With respect to the human body and its microbial inhabitants (the human microbiome), small-molecule-mediated interactions can define the difference between commensals and pathogens, and thus between health and disease states. We are interested in studying these interactions at three main levels: a) identifying the chemical repertoire of the human microbiome; b) characterizing the biological activities and functional roles of the identified molecules in mediating microbiome-host interactions in health and disease states; and finally c) using this knowledge to aid the diagnosis and treatment of human diseases via microbiome-targeted therapies.